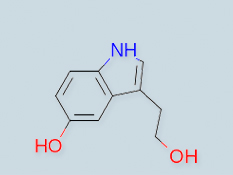
Automated pipeline for de novo metabolite identification using mass-spectrometry-based metabolomics
Metabolite identification is one of the biggest bottlenecks in metabolomics. Identifying human metabolites poses experimental, analytical, and computational challenges. Here we present a pipeline of previously developed cheminformatic tools and demonstrate how it facilitates metabolite identification using solely LC/MSn data. These tools process, annotate, and compare MSn data, and propose candidate structures for unknown metabolites either by identity assignment of identical mass spectral trees or by de novo identification using substructures of similar trees.